The Logic of DNA Replication Makes a Case for Intelligent Design
Why do I think God exists?
In short: The elegance, sophistication, and ingenuity of biochemical systems—and their astonishing similarity to man-made systems—convinces me that God is responsible for life’s origin and design.
While many skeptics readily acknowledge the remarkable designs of biochemical systems, they would disagree with my conclusion about God’s existence. Why? Because for every biochemical system I point to that displays beauty and elegance, they can point to one that seems to be poorly designed. In their view, these substandard designs reflect life’s evolutionary origin. They argue that evolutionary mechanisms kludged together the cell’s chemical systems through a historically contingent process that co-opted preexisting systems, cobbling them together to form new biochemical systems.
According to skeptics, one doesn’t have to look hard to find biochemical systems that seem to have been put together in a haphazard manner, and DNA replication appears to be an example of this. In many respects, DNA replication lies at the heart of the cell’s chemical operations. If designed by a Creator, this biochemical system, above all others, should epitomize intelligent design. Yet the DNA replication process appears to be unwieldy, inefficient, and unduly complex—the type of system evolution would generate by force, not the type of system worthy to be designated the product of the Creator’s handiwork.
Yet new work by Japanese researchers helps explain why DNA replication is the way it is.1 Instead of reflecting the cumbersome product of an unguided evolutionary history, the DNA replication process displays an exquisite molecular logic.
To appreciate the significance of the Japanese study and its implication for the creation/evolution controversy, a short biochemistry primer is in order. For readers who are familiar with DNA’s structure and the DNA replication process, you can skip the next two sections.
DNA
DNA consists of chain-like molecules known as polynucleotides. Two polynucleotide chains align in an antiparallel fashion to form a DNA molecule. (The two strands are arranged parallel to one another with the starting point of one strand in the polynucleotide duplex located next to the ending point of the other strand and vice versa.) The paired polynucleotide chains twist around each other to form the well-known DNA double helix. The cell’s machinery forms polynucleotide chains by linking together four different subunit molecules called nucleotides. The nucleotides used to build DNA chains are adenosine, guanosine, cytidine, and thymidine, famously abbreviated A, G, C, and T, respectively.
The nucleotide molecules that make up the strands of DNA are, in turn, complex molecules consisting of both a phosphate moiety, and a nucleobase (either adenine, guanine, cytosine, or thymine) joined to a 5-carbon sugar (deoxyribose).
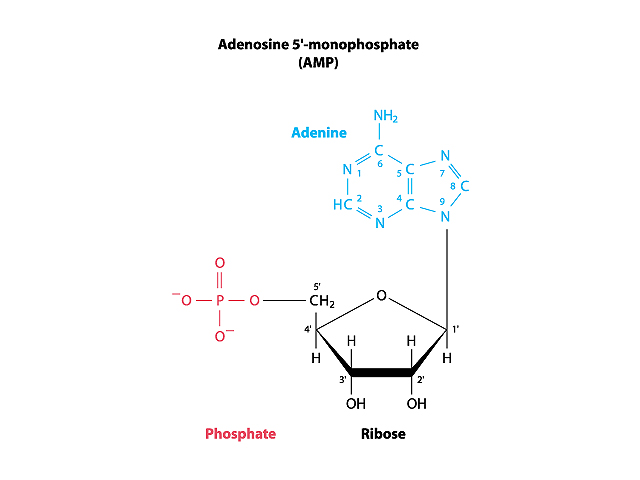
Image 1: Adenosine Monophosphate, a Nucleotide
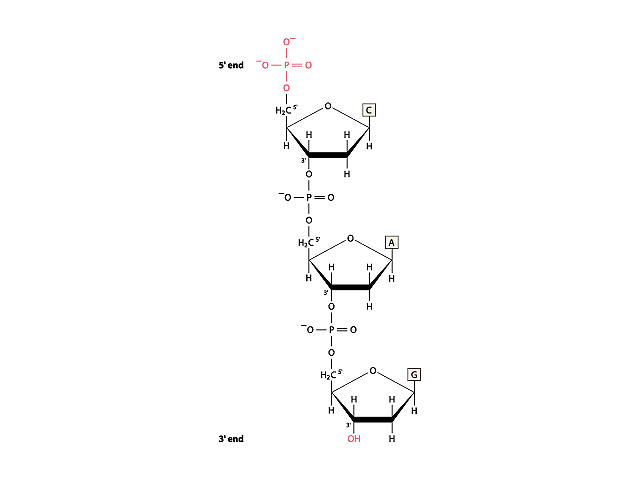
Repeatedly linking the phosphate group of one nucleotide to the deoxyribose unit of another nucleotide forms the backbone of the DNA strand. The nucleobases extend as side chains from the backbone of the DNA molecule and serve as interaction points when the two DNA strands align and twist to form the double helix.
Image 2: The DNA Backbone
When the two DNA strands align, the adenosine (A) side chains of one strand always pair with thymidine (T) side chains from the other strand. Likewise, the guanosine (G) side chains from one DNA strand always pair with cytidine (C) side chains from the other strand.
DNA Replication
Biochemists refer to DNA replication as a template-directed, semi-conservative process. By template-directed, biochemists mean that the nucleotide sequences of the “parent” DNA molecule function as a template, directing the assembly of the DNA strands of the two “daughter” molecules. By semi-conservative, biochemists mean that after replication, each daughter DNA molecule contains one newly formed DNA strand and one strand from the parent molecule.
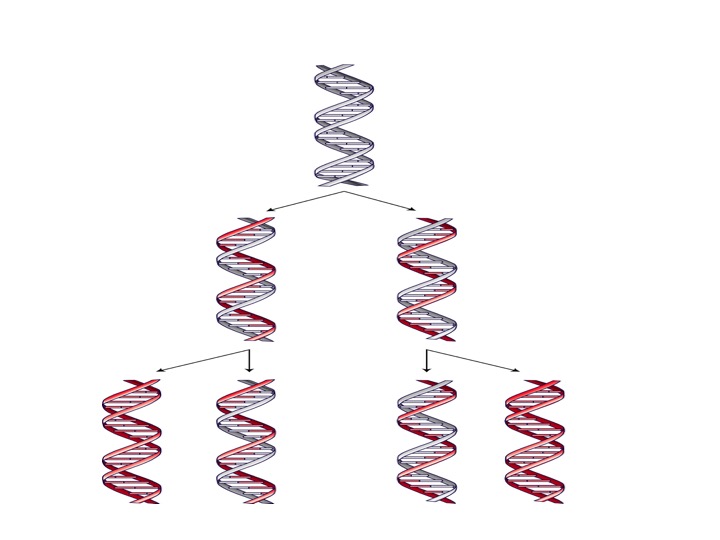
Image 3: Semi-Conservative DNA Replication
Conceptually, template-directed, semi-conservative DNA replication entails the separation of the parent DNA double-helix into two single strands. By using the base-pairing rules, each strand serves as a template for the cell’s machinery to use when it forms a new DNA strand with a nucleotide sequence complementary to the parent strand. Because each strand of the parent DNA molecule directs the production of a new DNA strand, two daughter molecules result. Each one possesses an original strand from the parent molecule and a newly formed DNA strand produced by a template-directed synthetic process.
DNA replication begins at specific sites along the DNA double helix, called replication origins. The DNA double helix unwinds locally at the origin of replication to produce what biochemists call a replication bubble. The bubble expands in both directions from the origin during the course of DNA replication. Once the individual strands of the DNA double helix unwind and are exposed within the replication bubble, they are available to direct the production of the daughter strand. The site where the DNA double helix continuously unwinds is called the replication fork. Because DNA replication proceeds in both directions away from the origin, there are two replication forks within each bubble.
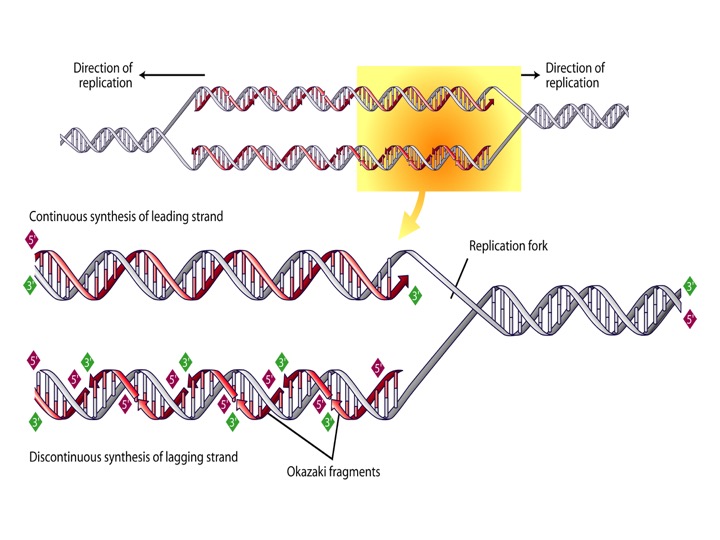
Image 4: DNA Replication
DNA replication can only proceed in a single direction, from the top of the DNA strand to the bottom. Because the strands that form the DNA double helix align in an antiparallel fashion with the top of one strand juxtaposed to the bottom of the other strand, only one strand at each replication fork has the proper orientation (bottom-to-top) to direct the assembly of a new strand, in the top-to-bottom direction. For this strand—referred to as the “leading strand”—DNA replication proceeds rapidly and continuously in the direction of the advancing replication fork.
DNA replication can’t proceed along the strand with the top-to-bottom orientation until the replication bubble has expanded enough to expose a sizable stretch of DNA. When this happens, DNA replication moves away from the advancing replication fork. DNA replication can only proceed a short distance for the top-to-bottom oriented strand before the replication process has to stop and wait for more of the parent DNA strand to be exposed. When a sufficient length of the parent DNA template is exposed for a second time, DNA replication can proceed again, but only briefly before it has to stop again and wait for more DNA to be exposed. The process of discontinuous DNA replication takes place repeatedly until the entire strand is replicated. Each time DNA replication starts and stops, a small fragment of DNA is produced. Biochemists refer to these pieces of DNA (that will eventually comprise the daughter strand) as “Okazaki fragments,” named after the biochemist who discovered them. Biochemists call the strand produced discontinuously the “lagging strand,” because DNA replication for this strand lags behind the more rapidly produced leading strand.
One additional point: The leading strand at one replication fork is the lagging strand at the other replication fork, since the replication forks at the two ends of the replication bubble advance in opposite directions.
Before the newly formed daughter strands can be produced, a small RNA primer must be produced. The protein that synthesizes new DNA by reading the parent DNA template strand—DNA polymerase—can’t start production from scratch. It has to be primed. A massive protein complex, called the primosome, which consists of more than 15 different proteins, produces the RNA primer needed by DNA polymerase.
Once primed, DNA polymerase will continuously produce DNA along the leading strand. However, for the lagging strand, DNA polymerase can only generate DNA in spurts to produce Okazaki fragments. Each time DNA polymerase generates an Okazaki fragment, the primosome complex must produce a new RNA primer.
Once DNA replication is completed, the RNA primers are removed from the continuous DNA of the leading strand and the Okazaki fragments that make up the lagging strand. A protein called a 3’–5’ exonuclease removes the RNA primers. A different DNA polymerase fills in the gaps created by the removal of the RNA primers. Finally, a protein called a ligase connects all the Okazaki fragments together to form a continuous piece of DNA out of the lagging strand.
DNA Replication and the Case for Evolution
This cursory description of DNA replication clearly illustrates the complexity of this biochemical operation. (Many details of the process were left out of the discussion.) This description also reveals why biochemists view this process as cumbersome and unwieldy. There is no obvious reason why DNA replication proceeds as a semi-conservative, RNA primer-dependent, unidirectional process involving leading and lagging strands to produce DNA daughter molecules. Because of this uncertainty, skeptics view DNA replication as a chance outcome of a historically contingent process, kludged together from the biochemical leftovers of the RNA world.
If there is one feature of DNA replication that is responsible for the complexity of the process, it is the directionality of DNA replication—from top to bottom. At first glance, it would seem as if the process would be simpler and more elegant if replication could proceed in both directions. Skeptics argue that the fact that it doesn’t reflects the evolutionary origin of the replication process.
Yet work by the team from Sapporo, Japan indicates that there is an exquisite molecular rationale for the directionality of DNA replication.
Why DNA Replication Proceeds in a Single Direction
These researchers recognized an important opportunity to ask why DNA replication proceeds only in a single direction with the discovery of a class of enzymes that add nucleotides to the ends of transfer RNA (tRNA) molecules. (tRNA molecules ferry amino acids to the ribsosome during protein synthesis.) If damaged, tRNA molecules cannot properly carry out their role in protein production. Fortunately, there are repair enzymes that can fix damaged tRNA molecules. One of them is called Thg-1-like protein (TLP).
TLP adds nucleotides to damaged ends of tRNA molecules. But instead of adding the nucleotides top to bottom, the enzyme adds these subunit molecules to the tRNA bottom to top, the opposite direction of DNA replication.
By determining the mechanism employed by TLP during bottom-to-top nucleotide addition, the researchers gained important insight into the constraints of DNA replication. As it turns out, bottom-to-top addition is a much more complex process than the normal top-to-bottom nucleotide addition. Bottom-to-top addition is a cumbersome two-step process that requires an enzyme with two active sites that have to be linked together in a precise way. In contrast, top-to-bottom addition is a simple, one-step reaction that proceeds with a single active site. In other words, DNA replication proceeds in a single direction (top-to-bottom) because it is mechanistically simpler and more efficient.
One could argue that the complexity that arises by the top-to-bottom DNA replication process is a trade-off for a mechanistically simpler nucleotide addition reaction. Still, if DNA replication proceeded in both directions the process would be complex and unwieldy. For example, if replication proceeded in two directions, the cell would require two distinct types of primosomes and DNA polymerases, one set for each direction of DNA replication. Employing two sets of primosomes and DNA polymerases is clearly less efficient than employing a single set of enzymes.
Ironically, if DNA replication could proceed in two directions, there still would be a leading and a lagging strand. Why? Because bottom-to-top replication is a two-step process and would proceed more slowly than the single step of top-to-bottom replication. In other words, the assembly of the DNA strand in a bottom-to-top direction would lag behind the assembly of the DNA strand that traveled in a top-to-bottom direction.
Bidirectional DNA replication would also cause another complication due to a crowding effect. Once the replication bubble opens, both sets of replication enzymes would have to fit into the replication bubble’s constrained space. This molecular overcrowding would further compromise the efficiency of the replication process. Overcrowding is not an issue for unidirectional DNA replication that proceeds in a top-to-bottom direction.
The bottom line: In light of this new insight, it is hard to argue that DNA replication has been cobbled together via a historically contingent pathway. Instead, it is looking more and more like a process ingenuously designed by a Divine Mind.
Resources
The Cell’s Design: How Chemistry Reveals the Creator’s Artistry by Fazale Rana (book)
“DNA Soaks Up Sun’s Rays” by Fazale Rana (article)
“DNA: Designed for Flexibility” by Fazale Rana (article)
“How the Central Dogma of Molecular Biology Points to Design” by Fazale Rana (article)
“Why I Believe God Exists: Evidences from a Biochemist” by Fazale Rana (video)
Endnotes
- Shoko Kimura et al., “Template-Dependent Nucleotide Addition in the Reverse (3’–5’) Direction by Thg1-like Protein,” Science Advances 2 (March 2016): e1501397, doi:10.1126/sciadv.1501397.